Introduction
This article is a brief illustration of how main functions from the package manymome (Cheung & Cheung, 2024) can be used in some typical cases. It assumes that readers have basic understanding of mediation, moderation, moderated mediation, structural equation modeling (SEM), and bootstrapping.
Workflow
The use of manymome adopts a two-stage workflow:
-
Stage 1: Fit the model
This can be done by SEM (using
lavaan::sem()) or a series of regression (usinglm()).When
lavaan::sem()is used, no need to label any parameters or denote any variables as the predictors, mediators, moderators, or outcome variables for computing indirect effects or conditional indirect effects. Stage 2 will take care of this.
-
Stage 2: Compute the indirect effects and conditional indirect effects
This can be done along nearly any path in the model for any levels of the moderators.
Just specify the start (
x), the mediator(s) (m, if any), and the end (y) for indirect effects. The functions will find the coefficients automatically.If a path has one or more moderators, conditional indirect effects can be computed. Product terms will be identified automatically.
The levels of the moderators can be decided in this stage and can be changed as often as needed.
Bootstrapping confidence intervals: All main
functions support bootstrap confidence intervals for the effects.
Bootstrapping can done in Stage 1 (e.g., by lavaan::sem()
using se = "boot") or in Stage 2 in the first call to the
main functions, and only needs to be conducted once. Alternatively,
do_boot() can be use (see
vignette("do_boot")). The bootstrap estimates can be reused
by most main functions of manymome for any path and any
level of the moderators.
Monte Carlo confidence intervals: Initial support
for Monte Carlo confidence interval has been added to all main functions
for the effects in a model fitted by lavaan::sem(). The
recommended workflow is to use do_mc() to generate the
simulated sampling estimates. The simulated estimates can be reused by
most main functions of manymome for any path and any level
of the moderators. To keep the length of this vignette short, it only
covers bootstrapping confidence intervals. Please see
vignette("do_mc") for an illustration on how to form Monte
Carlo confidence intervals.
Standardized effects: All main functions in Stage 2 support standardized effects and form their bootstrap confidence interval correctly (Cheung, 2009; Friedrich, 1982). No need to standardize the variables in advance in Stage 1, even for paths with moderators.
What Will Be Covered In This Get-Started Article
Use
cond_indirect_effects()to compute conditional indirect effects, with bootstrap confidence intervals.Use
indirect_effect()to compute an indirect effect, with bootstrap confidence interval.Use
+and-to compute a function of effects, such as total indirect effects or total effects.Use
do_boot()to generate bootstrap estimates forcond_indirect_effects(),indirect_effect(), and some other functions inmanymome.Use
index_of_mome()andindex_of_momome()to compute the index of moderated mediation and the index of moderated moderated mediation, respectively, with bootstrap confidence intervals.Compute standardized conditional indirect effects and standardized indirect effect using
cond_indirect_effects()andindirect_effect(), respectively.
Moderated Mediation by SEM using lavaan
This is the sample data set comes with the package:
library(manymome)
dat <- data_med_mod_ab
print(head(dat), digits = 3)
#> x w1 w2 m y c1 c2
#> 1 9.27 4.97 2.66 3.46 8.80 9.26 3.14
#> 2 10.79 4.13 3.33 4.05 7.37 10.71 5.80
#> 3 11.10 5.91 3.32 4.04 8.24 10.60 5.45
#> 4 9.53 4.78 2.32 3.54 8.37 9.22 3.83
#> 5 10.00 4.38 2.95 4.65 8.39 9.58 4.26
#> 6 12.25 5.81 4.04 4.73 9.65 9.51 4.01Suppose this is the model being fitted:
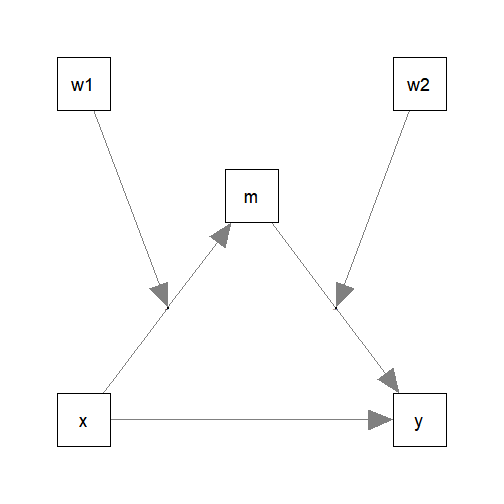
The models are intended to be simple enough for illustration but
complicated enough to show the flexibility of manymome.
More complicated models are also supported, discussed later.
Fitting the Model
The model fitted above is a moderated mediation model with
a mediation path
x -> m -> y, and-
two moderators:
x -> mmoderated byw1m -> ymoderated byw2.
The effects of interest are the conditional indirect effects: the indirect effects
from
xtoythrough
mfor different levels of
w1andw2.
cond_indirect_effects() can estimate these effects in
the model fitted by lavaan::sem(). There is no need to
label any path coefficients or define any user parameters (but users
can, if so desired; they have no impact on the functions in
manymome). To illustrate a more realistic scenario, two
control variables, c1 and c2, are also
included.
library(lavaan)
# Form the product terms
dat$w1x <- dat$w1 * dat$x
dat$w2m <- dat$w2 * dat$m
mod <-
"
m ~ x + w1 + w1x + c1 + c2
y ~ m + w2 + w2m + x + c1 + c2
# Covariances of the error term of m with w2m and w2
m ~~ w2m + w2
# Covariance between other variables
# They need to be added due to the covariances added above
# See Kwan and Chan (2018) and Miles et al. (2015)
w2m ~~ w2 + x + w1 + w1x + c1 + c2
w2 ~~ x + w1 + w1x + c1 + c2
x ~~ w1 + w1x + c1 + c2
w1 ~~ w1x + c1 + c2
w1x ~~ c1 + c2
c1 ~~ c2
"
fit <- sem(model = mod,
data = dat,
fixed.x = FALSE,
estimator = "MLR")MLR is used to take into account probable nonnormality
due to the product terms. fixed.x = FALSE is used to allow
the predictors to be random variables. This is usually necessary when
the values of the predictor are also sampled from the populations, and
so their standard deviations are sample statistics.
These are the parameter estimates of the paths:
parameterEstimates(fit)[parameterEstimates(fit)$op == "~", ]
#> lhs op rhs est se z pvalue ci.lower ci.upper
#> 1 m ~ x -0.663 0.533 -1.244 0.213 -1.707 0.381
#> 2 m ~ w1 -2.290 1.010 -2.267 0.023 -4.269 -0.310
#> 3 m ~ w1x 0.204 0.101 2.023 0.043 0.006 0.401
#> 4 m ~ c1 -0.020 0.079 -0.251 0.801 -0.175 0.135
#> 5 m ~ c2 -0.130 0.090 -1.444 0.149 -0.306 0.046
#> 6 y ~ m -0.153 0.248 -0.616 0.538 -0.638 0.333
#> 7 y ~ w2 -0.921 0.401 -2.300 0.021 -1.706 -0.136
#> 8 y ~ w2m 0.204 0.079 2.579 0.010 0.049 0.359
#> 9 y ~ x 0.056 0.086 0.653 0.514 -0.113 0.225
#> 10 y ~ c1 -0.102 0.081 -1.261 0.207 -0.261 0.056
#> 11 y ~ c2 -0.108 0.087 -1.249 0.212 -0.279 0.062The moderation effects of both w1 and w2
are significant. The indirect effect from x to
y through m depends on the level of
w1 and w2.
Conditional Indirect Effects
To form bootstrap confidence intervals, bootstrapping needs to be
done. There are several ways to do this. We first illustrate using
do_boot().
Do Bootstrapping (Once)
Using do_boot() instead of setting se to
"boot" when calling lavaan::sem() allows users
to use other method for standard errors and confidence intervals for
other parameters, such as the various types of robust standard errors
provided by lavaan::sem().
The function do_boot() is used to generate and store
bootstrap estimates as well as implied variances of variables, which are
needed to estimate standardized effects.
fit_boot <- do_boot(fit = fit,
R = 500,
seed = 53253,
ncores = 1)These are the major arguments:
fit: The output oflavaan::sem().R: The number of bootstrap samples, which should be 2000 or even 5000 in real research.Ris set to 500 here just for illustration.seed: The seed to reproduce the results.ncores: The number of processes in parallel processing. The default number is the number of detected physical cores minus 1. Can be omitted in real studies. Set to 1 here for illustration.
By default, parallel processing is used, and so the results are
reproducible with the same seed only if the number of processes is the
same. See do_boot() for other options and
vignette("do_boot") on the output of
do_boot().
The output, fit_boot in this case, can then be used for
all subsequent analyses on this model.
Estimate Conditional Indirect Effects
To compute conditional indirect effects and form bootstrap confidence
intervals, we can use cond_indirect_effects().
out_cond <- cond_indirect_effects(wlevels =c("w1", "w2"),
x = "x",
y = "y",
m = "m",
fit = fit,
boot_ci = TRUE,
boot_out = fit_boot)These are the major arguments:
-
wlevels: The vector of the names of the moderators. Order does not matter. If the default levels are not suitable, custom levels can be created by functions likemod_levels()andmerge_mod_levels()(seevignette("mod_levels")). -
x: The name of the predictor. -
y: The name of the outcome variable. -
m: The name of the mediator, or a vector of names if the path has more than one mediator (see this example). -
fit: The output oflavaan::sem(). -
boot_ci: Set toTRUEto request bootstrap confidence intervals. Default isFALSE. -
boot_out: The pregenerated bootstrap estimates generated bydo_boot()or previous call tocond_indirect_effects()orindirect_effect().
This is the output:
out_cond
#>
#> == Conditional indirect effects ==
#>
#> Path: x -> m -> y
#> Conditional on moderator(s): w1, w2
#> Moderator(s) represented by: w1, w2
#>
#> [w1] [w2] (w1) (w2) ind CI.lo CI.hi Sig m~x y~m
#> 1 M+1.0SD M+1.0SD 6.173 4.040 0.399 0.139 0.705 Sig 0.596 0.671
#> 2 M+1.0SD M-1.0SD 6.173 2.055 0.158 -0.025 0.381 0.596 0.266
#> 3 M-1.0SD M+1.0SD 4.038 4.040 0.107 -0.148 0.358 0.160 0.671
#> 4 M-1.0SD M-1.0SD 4.038 2.055 0.043 -0.062 0.191 0.160 0.266
#>
#> - [CI.lo to CI.hi] are 95.0% percentile confidence intervals by
#> nonparametric bootstrapping with 500 samples.
#> - The 'ind' column shows the conditional indirect effects.
#> - 'm~x','y~m' is/are the path coefficient(s) along the path conditional
#> on the moderator(s).For two or more moderators, the default levels for numeric moderators are one standard deviation (SD) below mean and one SD above mean. For two moderators, there are four combinations.
As shown above, among these four sets of levels, the indirect effect
from x to y through m is
significant only when both w1 and w2 are one
SD above their means. The indirect effect at these levels of
w1 and w2 are 0.399, with 95% bootstrap
confidence interval [0.139, 0.705].
The function cond_indirect_effects(), as well as other
functions described below, also supports bias-corrected (BC) confidence
interval, which can be requested by adding boot_type = "bc"
to the call. However, authors in some recent work do not advocate this
method (e.g., Falk & Biesanz, 2015; Hayes, 2022; Tofighi &
Kelley, 2020). Therefore, this option is provided merely for research
purpose.
Examine the Effect at a Particular Set of Levels of the Moderators
To learn more about the conditional effect for one combination of the
levels of the moderators, get_one_cond_indirect_effect()
can be used, with the first argument the output of
cond_indirect_effects() and the second argument the row
number. For example, this shows the details on the computation of the
indirect effect when both w1 and w2 are one SD
above their means (row 1):
get_one_cond_indirect_effect(out_cond, 1)
#>
#> == Conditional Indirect Effect ==
#>
#> Path: x -> m -> y
#> Moderators: w1, w2
#> Conditional Indirect Effect: 0.399
#> 95.0% Bootstrap CI: [0.139 to 0.705]
#> When: w1 = 6.173, w2 = 4.040
#>
#> Computation Formula:
#> (b.m~x + (b.w1x)*(w1))*(b.y~m + (b.w2m)*(w2))
#>
#> Computation:
#> ((-0.66304) + (0.20389)*(6.17316))*((-0.15271) + (0.20376)*(4.04049))
#>
#>
#> Percentile confidence interval formed by nonparametric bootstrapping
#> with 500 bootstrap samples.
#>
#> Coefficients of Component Paths:
#> Path Conditional Effect Original Coefficient
#> m~x 0.596 -0.663
#> y~m 0.671 -0.153Changing the Levels of the Moderators
The levels of the moderators, w1 and w2 in
this example, can be controlled directly by users. For examples,
percentiles or exact values of the moderators can be used. See
vignette("mod_levels") on how to specify other levels of
the moderators, and the arguments w_method,
sd_from_mean, and percentiles of
cond_indirect_effects().
Standardized Conditional Indirect Effects
To compute the standardized conditional indirect effects, we can
standardize only the predictor (x), only the outcome
(y), or both.
To standardize x, set standardized_x to
TRUE. To standardize y, set
standardized_y to TRUE. To standardize both,
set both standardized_x and standardized_y to
TRUE.
This is the result when both x and y are
standardized:
out_cond_stdxy <- cond_indirect_effects(wlevels =c("w1", "w2"),
x = "x",
y = "y",
m = "m",
fit = fit,
boot_ci = TRUE,
boot_out = fit_boot,
standardized_x = TRUE,
standardized_y = TRUE)Note that fit_boot is used so that there is no need to
do bootstrapping again.
This is the output:
out_cond_stdxy
#>
#> == Conditional indirect effects ==
#>
#> Path: x -> m -> y
#> Conditional on moderator(s): w1, w2
#> Moderator(s) represented by: w1, w2
#>
#> [w1] [w2] (w1) (w2) std CI.lo CI.hi Sig m~x y~m ind
#> 1 M+1.0SD M+1.0SD 6.173 4.040 0.401 0.154 0.655 Sig 0.596 0.671 0.399
#> 2 M+1.0SD M-1.0SD 6.173 2.055 0.159 -0.029 0.363 0.596 0.266 0.158
#> 3 M-1.0SD M+1.0SD 4.038 4.040 0.108 -0.145 0.370 0.160 0.671 0.107
#> 4 M-1.0SD M-1.0SD 4.038 2.055 0.043 -0.062 0.190 0.160 0.266 0.043
#>
#> - [CI.lo to CI.hi] are 95.0% percentile confidence intervals by
#> nonparametric bootstrapping with 500 samples.
#> - std: The standardized conditional indirect effects.
#> - ind: The unstandardized conditional indirect effects.
#> - 'm~x','y~m' is/are the path coefficient(s) along the path conditional
#> on the moderator(s).The standardized indirect effect when both w1 and
w2 are one SD above mean is 0.401, with 95% bootstrap
confidence interval [0.154, 0.655].
That is, when both w1 and w2 are one SD
above their means, if x increases by one SD, it leads to an
increase of 0.401 SD of y through m.
Index of Moderated Moderated Mediation
The index of moderated moderated mediation (Hayes, 2018) can be
estimated, along with bootstrap confidence interval, using the function
index_of_momome():
out_momome <- index_of_momome(x = "x",
y = "y",
m = "m",
w = "w1",
z = "w2",
fit = fit,
boot_ci = TRUE,
boot_out = fit_boot)These are the major arguments:
-
x: The name of the predictor. -
y: The name of the outcome variable. -
m: The name of the mediator, or a vector of names if the path has more than one mediator (see this example). -
w: The name of one of the moderator. -
z: The name of the other moderator. The order ofwandzdoes not matter. -
fit: The output oflavaan::sem(). -
boot_ci: Set toTRUEto request bootstrap confidence intervals. Default isFALSE. -
boot_out: The pregenerated bootstrap estimates generated bydo_boot()or previous call tocond_indirect_effects()andindirect_effect().
This is the result:
out_momome
#>
#> == Conditional indirect effects ==
#>
#> Path: x -> m -> y
#> Conditional on moderator(s): w1, w2
#> Moderator(s) represented by: w1, w2
#>
#> [w1] [w2] (w1) (w2) ind CI.lo CI.hi Sig m~x y~m
#> 1 1 1 1 1 -0.023 -0.276 0.312 -0.459 0.051
#> 2 1 0 1 0 0.070 -0.206 0.649 -0.459 -0.153
#> 3 0 1 0 1 -0.034 -0.364 0.383 -0.663 0.051
#> 4 0 0 0 0 0.101 -0.252 0.868 -0.663 -0.153
#>
#> == Index of Moderated Moderated Mediation ==
#>
#> Levels compared:
#> (Row 1 - Row 2) - (Row 3 - Row 4)
#>
#> x y Index CI.lo CI.hi
#> Index x y 0.042 -0.003 0.116
#>
#> - [CI.lo, CI.hi]: 95% percentile confidence interval.The index of moderated moderated mediation is 0.042, with 95% bootstrap confidence interval [-0.003, 0.116].
Note that this index is specifically for the change when
w1 or w2 increases by one unit.
Index of Moderated Mediation
The manymome package also has a function to compute the
index of moderated mediation (Hayes, 2015). Suppose we modify
the model and remove one of the moderators:
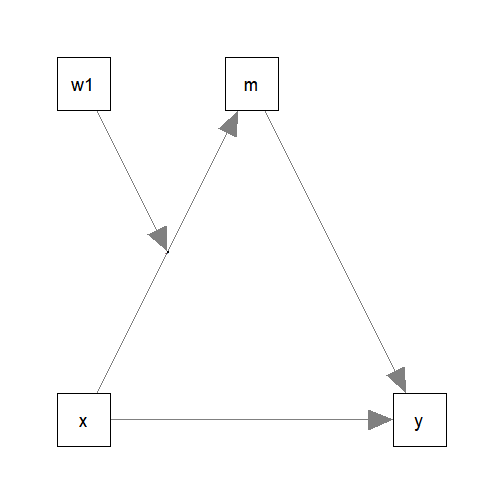
This is the lavaan model:
library(lavaan)
dat$w1x <- dat$w1 * dat$x
mod2 <-
"
m ~ x + w1 + w1x + c1 + c2
y ~ m + x + c1 + c2
"
fit2 <- sem(model = mod2,
data = dat,
fixed.x = FALSE,
estimator = "MLR")These are the parameter estimates of the paths:
parameterEstimates(fit2)[parameterEstimates(fit2)$op == "~", ]
#> lhs op rhs est se z pvalue ci.lower ci.upper
#> 1 m ~ x -0.663 0.533 -1.244 0.213 -1.707 0.381
#> 2 m ~ w1 -2.290 1.010 -2.267 0.023 -4.269 -0.310
#> 3 m ~ w1x 0.204 0.101 2.023 0.043 0.006 0.401
#> 4 m ~ c1 -0.020 0.079 -0.251 0.801 -0.175 0.135
#> 5 m ~ c2 -0.130 0.090 -1.444 0.149 -0.306 0.046
#> 6 y ~ m 0.434 0.114 3.815 0.000 0.211 0.657
#> 7 y ~ x 0.053 0.093 0.570 0.569 -0.130 0.237
#> 8 y ~ c1 -0.108 0.080 -1.352 0.177 -0.265 0.049
#> 9 y ~ c2 -0.077 0.085 -0.904 0.366 -0.243 0.090We generate the bootstrap estimates first (R should be
2000 or even 5000 in real research):
fit2_boot <- do_boot(fit = fit2,
R = 500,
seed = 53253,
ncores = 1)The function index_of_mome() can be used to compute the
index of moderated mediation of w1 on the path
x -> m -> y:
out_mome <- index_of_mome(x = "x",
y = "y",
m = "m",
w = "w1",
fit = fit2,
boot_ci = TRUE,
boot_out = fit2_boot)The arguments are nearly identical to those of
index_of_momome(), except that only w needs to
be specified. This is the output:
out_mome
#>
#> == Conditional indirect effects ==
#>
#> Path: x -> m -> y
#> Conditional on moderator(s): w1
#> Moderator(s) represented by: w1
#>
#> [w1] (w1) ind CI.lo CI.hi Sig m~x y~m
#> 1 1 1 -0.199 -0.762 0.230 -0.459 0.434
#> 2 0 0 -0.288 -0.998 0.222 -0.663 0.434
#>
#> == Index of Moderated Mediation ==
#>
#> Levels compared: Row 1 - Row 2
#>
#> x y Index CI.lo CI.hi
#> Index x y 0.088 -0.006 0.223
#>
#> - [CI.lo, CI.hi]: 95% percentile confidence interval.In this model, the index of moderated mediation is 0.088, with 95%
bootstrap confidence interval [-0.006, 0.223]. The indirect effect of
x on y through m does not
significantly change when w1 increases by one unit.
Note that this index is specifically for the change when
w1 increases by one unit. The index being not significant
does not contradict with the significant moderation effect suggested by
the product term.
Mediation Only
The package can also be used for a mediation model.
This is the sample data set that comes with the package:
library(manymome)
dat <- data_serial
print(head(dat), digits = 3)
#> x m1 m2 y c1 c2
#> 1 12.12 20.6 9.33 9.00 0.109262 6.01
#> 2 9.81 18.2 9.47 11.56 -0.124014 6.42
#> 3 10.11 20.3 10.05 9.35 4.278608 5.34
#> 4 10.07 19.7 10.17 11.41 1.245356 5.59
#> 5 11.91 20.5 10.05 14.26 -0.000932 5.34
#> 6 9.13 16.5 8.93 10.01 1.802727 5.91Suppose this is the model being fitted, with c1 and
c2 the control variables.
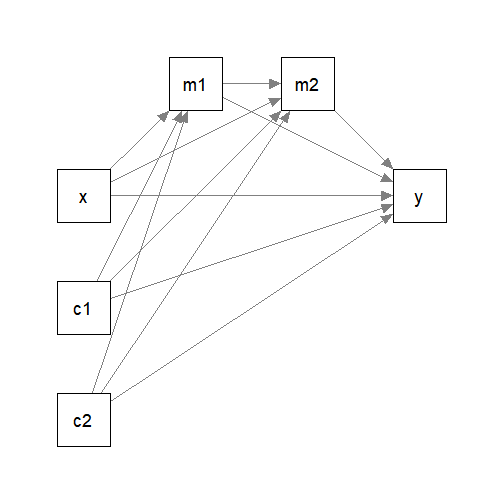
Fitting the Model
Fitting this model in lavaan::sem() is very simple. With
manymome, there is no need to label paths or define user
parameters for the indirect effects.
mod_med <- "
m1 ~ x + c1 + c2
m2 ~ m1 + x + c1 + c2
y ~ m2 + m1 + x + c1 + c2
"
fit_med <- sem(model = mod_med,
data = dat,
fixed.x = TRUE)These are the estimates of the paths:
parameterEstimates(fit_med)[parameterEstimates(fit_med)$op == "~", ]
#> lhs op rhs est se z pvalue ci.lower ci.upper
#> 1 m1 ~ x 0.822 0.092 8.907 0.000 0.641 1.003
#> 2 m1 ~ c1 0.171 0.089 1.930 0.054 -0.003 0.346
#> 3 m1 ~ c2 -0.189 0.091 -2.078 0.038 -0.367 -0.011
#> 4 m2 ~ m1 0.421 0.099 4.237 0.000 0.226 0.615
#> 5 m2 ~ x -0.116 0.123 -0.946 0.344 -0.357 0.125
#> 6 m2 ~ c1 0.278 0.090 3.088 0.002 0.101 0.454
#> 7 m2 ~ c2 -0.162 0.092 -1.756 0.079 -0.343 0.019
#> 8 y ~ m2 0.521 0.221 2.361 0.018 0.088 0.953
#> 9 y ~ m1 -0.435 0.238 -1.830 0.067 -0.902 0.031
#> 10 y ~ x 0.493 0.272 1.811 0.070 -0.040 1.026
#> 11 y ~ c1 0.099 0.208 0.476 0.634 -0.308 0.506
#> 12 y ~ c2 -0.096 0.207 -0.465 0.642 -0.501 0.309Estimate Indirect Effects
indirect_effect() can be used to estimate an indirect
effect and form its bootstrapping confidence interval along a path in a
model that starts with any numeric variable, ends with any numeric
variable, through any numeric variable(s).
We illustrate another approach to generate bootstrap estimates: using
indirect_effect() to do both bootstrapping and estimate the
indirect effect.
For example, this is the call for the indirect effect from
x to y through m1 and
m2:
out_med <- indirect_effect(x = "x",
y = "y",
m = c("m1", "m2"),
fit = fit_med,
boot_ci = TRUE,
R = 500,
seed = 43143,
ncores = 1)The main arguments are:
-
x: The name of the predictor. The start of the path. -
y: The name of the outcome variable. The end of the path. -
m: The name of the mediator, or the vector of names of the mediators if the path has more than one mediator, as in this example. The path moves from the first mediator to the last mediator. In this example, the correct order isc("m1", "m2"). -
fit: The output oflavaan::sem(). -
boot_ci: Set toTRUEto request bootstrapping confidence intervals. Default isFALSE. -
R: The number of bootstrap samples. Only 500 bootstrap samples for illustration. SetRto 2000 or even 5000 in real research. -
seed: The seed for the random number generator. -
ncores: The number of processes in parallel processing. The default number is the number of detected physical cores minus 1. Can be omitted in real studies. Set to 1 here for illustration.
Like do_boot(), by default, parallel processing is used,
and so the results are reproducible with the same seed only if the
number of processes (cores) is the same.
This is the output:
out_med
#>
#> == Indirect Effect ==
#>
#> Path: x -> m1 -> m2 -> y
#> Indirect Effect: 0.180
#> 95.0% Bootstrap CI: [0.034 to 0.396]
#>
#> Computation Formula:
#> (b.m1~x)*(b.m2~m1)*(b.y~m2)
#>
#> Computation:
#> (0.82244)*(0.42078)*(0.52077)
#>
#>
#> Percentile confidence interval formed by nonparametric bootstrapping
#> with 500 bootstrap samples.
#>
#> Coefficients of Component Paths:
#> Path Coefficient
#> m1~x 0.822
#> m2~m1 0.421
#> y~m2 0.521The indirect effect from x to y through
m1 and m2 is 0.180, with a 95% confidence
interval of [0.034, 0.396], significantly different from zero
(p < .05).
Because bootstrap confidence interval is requested, the bootstrap
estimates are stored in out_med. This output from
indirect_effect() can also be used in the argument
boot_out of other functions.
Standardized Indirect Effect
To compute the indirect effect with the predictor standardized, set
standardized_x to TRUE. To compute the
indirect effect with the outcome variable standardized, set
standardized_y to TRUE. To compute the
(completely) standardized indirect effect, set both
standardized_x and standardized_y to
TRUE.
This is the call to compute the (completely) standardized indirect effect:
out_med_stdxy <- indirect_effect(x = "x",
y = "y",
m = c("m1", "m2"),
fit = fit_med,
boot_ci = TRUE,
boot_out = out_med,
standardized_x = TRUE,
standardized_y = TRUE)
out_med_stdxy
#>
#> == Indirect Effect (Both 'x' and 'y' Standardized) ==
#>
#> Path: x -> m1 -> m2 -> y
#> Indirect Effect: 0.086
#> 95.0% Bootstrap CI: [0.017 to 0.183]
#>
#> Computation Formula:
#> (b.m1~x)*(b.m2~m1)*(b.y~m2)*sd_x/sd_y
#>
#> Computation:
#> (0.82244)*(0.42078)*(0.52077)*(0.95010)/(1.99960)
#>
#>
#> Percentile confidence interval formed by nonparametric bootstrapping
#> with 500 bootstrap samples.
#>
#> Coefficients of Component Paths:
#> Path Coefficient
#> m1~x 0.822
#> m2~m1 0.421
#> y~m2 0.521
#>
#> NOTE:
#> - The effects of the component paths are from the model, not
#> standardized.The indirect effect from x to y through
m1 and m2 is 0.086, with a 95% confidence
interval of [0.017, 0.183], significantly different from zero
(p < .05). One SD increase of x leads to 0.086
increase in SD of y through m1 and
m2.
Estimating Indirect Effects For Any Paths
indirect_effect() can be used for the indirect effect in
any path in a path model.
For example, to estimate and test the indirect effect from
x through m2 to y, bypassing
m1, simply set x to "x",
y to "y", and m to
"m2":
out_x_m2_y <- indirect_effect(x = "x",
y = "y",
m = "m2",
fit = fit_med,
boot_ci = TRUE,
boot_out = out_med)
out_x_m2_y
#>
#> == Indirect Effect ==
#>
#> Path: x -> m2 -> y
#> Indirect Effect: -0.060
#> 95.0% Bootstrap CI: [-0.232 to 0.097]
#>
#> Computation Formula:
#> (b.m2~x)*(b.y~m2)
#>
#> Computation:
#> (-0.11610)*(0.52077)
#>
#>
#> Percentile confidence interval formed by nonparametric bootstrapping
#> with 500 bootstrap samples.
#>
#> Coefficients of Component Paths:
#> Path Coefficient
#> m2~x -0.116
#> y~m2 0.521The indirect effect along this path is not significant.
Similarly, indirect effects from m1 through
m2 to y or from x through
m1 to y can also be tested by setting the
three arguments accordingly. Although c1 and
c2 are labelled as control variables, if appropriate, their
indirect effects on y through m1 and/or
m2 can also be computed and tested.
Total Indirect Effects and Total Effects
Addition (+) and subtraction (-) can be
applied to the outputs of indirect_effect(). For example,
the total indirect effect from x to y
is the sum of these indirect effects:
x -> m1 -> m2 -> yx -> m1 -> yx -> m2 -> y
Two of them have been computed above (out_med and
out_x_m2_y). We compute the indirect effect in
x -> m1 -> y
out_x_m1_y <- indirect_effect(x = "x",
y = "y",
m = "m1",
fit = fit_med,
boot_ci = TRUE,
boot_out = out_med)
out_x_m1_y
#>
#> == Indirect Effect ==
#>
#> Path: x -> m1 -> y
#> Indirect Effect: -0.358
#> 95.0% Bootstrap CI: [-0.747 to -0.017]
#>
#> Computation Formula:
#> (b.m1~x)*(b.y~m1)
#>
#> Computation:
#> (0.82244)*(-0.43534)
#>
#>
#> Percentile confidence interval formed by nonparametric bootstrapping
#> with 500 bootstrap samples.
#>
#> Coefficients of Component Paths:
#> Path Coefficient
#> m1~x 0.822
#> y~m1 -0.435We can then “add” the indirect effects to get the total indirect effect:
total_ind <- out_med + out_x_m1_y + out_x_m2_y
total_ind
#>
#> == Indirect Effect ==
#>
#> Path: x -> m1 -> m2 -> y
#> Path: x -> m1 -> y
#> Path: x -> m2 -> y
#> Function of Effects: -0.238
#> 95.0% Bootstrap CI: [-0.596 to 0.092]
#>
#> Computation of the Function of Effects:
#> ((x->m1->m2->y)
#> +(x->m1->y))
#> +(x->m2->y)
#>
#>
#> Percentile confidence interval formed by nonparametric bootstrapping
#> with 500 bootstrap samples.The total indirect effect is -0.238, not significant. This is an example of inconsistent mediation: some of the indirect Effects are positive and some are negative:
coef(out_med)
#> y~x
#> 0.1802238
coef(out_x_m1_y)
#> y~x
#> -0.3580391
coef(out_x_m2_y)
#> y~x
#> -0.060461Similarly, the total effect of x on y can
be computed by adding all the effects, direct or indirect. The direct
effect can be computed with m not set:
out_x_direct <- indirect_effect(x = "x",
y = "y",
fit = fit_med,
boot_ci = TRUE,
boot_out = out_med)
out_x_direct
#>
#> == Effect ==
#>
#> Path: x -> y
#> Effect: 0.493
#> 95.0% Bootstrap CI: [-0.041 to 1.045]
#>
#> Computation Formula:
#> (b.y~x)
#>
#> Computation:
#> (0.49285)
#>
#>
#> Percentile confidence interval formed by nonparametric bootstrapping
#> with 500 bootstrap samples.This is the total effect:
total_effect <- out_med + out_x_m1_y + out_x_m2_y + out_x_direct
total_effect
#>
#> == Indirect Effect ==
#>
#> Path: x -> m1 -> m2 -> y
#> Path: x -> m1 -> y
#> Path: x -> m2 -> y
#> Path: x -> y
#> Function of Effects: 0.255
#> 95.0% Bootstrap CI: [-0.200 to 0.731]
#>
#> Computation of the Function of Effects:
#> (((x->m1->m2->y)
#> +(x->m1->y))
#> +(x->m2->y))
#> +(x->y)
#>
#>
#> Percentile confidence interval formed by nonparametric bootstrapping
#> with 500 bootstrap samples.The total effect is 0.255, not significant. This illustrates that total effect can be misleading when the component paths are of different signs.
See help(math_indirect) for more information of addition
and subtraction for the output of indirect_effect().
Summary
Advantages
The model fitting stage is easier. No need to label any parameters or define any effects. Users can also use other methods for confidence interval and use bootstrapping only for indirect effects and conditional indirect effects.
Missing data can be be handled by
missing = "fiml"in callinglavaan::sem(). Because bootstrapping estimates are used in Stage 2, indirect effects and conditional indirect effects can also be computed with bootstrap confidence intervals, just like defining them inlavaan, in the presence of missing data.Missing data handled by multiple imputation is also supported since version 0.2.7.3. Models fitted by
lavaan.mi::sem.mi()orlavaan.mi::lavaan.mi()to multiple imputation datasets can be used just like the output oflavaan::sem(). Monte Carlo confidence intervals cna be formed for effects computed for these models (seevignette("do_mc_lavaan_mi")). (Support forsemTools::runMI()and its wrappers was removed in version 0.2.7.3 because this and related functions were deprecated insemTools.)Bootstrapping only needs to be done once. The bootstrap estimates can be reused in computing indirect effects and conditional indirect effects. This is particularly useful when the sample size is large and there is missing data.
Users can explore any path for any levels of the moderators without respecifying and refitting the model.
Flexibility makes it difficult to test all possible scenarios. Therefore, the print methods will also print the details of the computation (e.g., how an indirect effect is computed) so that users can (a) understand how each effect is computed, and (b) verify the computation if necessary.
See this section for other advantages.
Limitations
The package manymome supports “many” models … but
certainly not all. There are models that it does not yet support. For
example, it does not support a path that starts with a nominal
categorical variable (except for a dichotomous variable). Other tools
need to be used for these cases. See this
section for other limitations.
Other Uses and Scenarios
There are other options available in manymome. For
example, it can be used for categorical moderators and models fitted by
multiple regression. Please refer to the help page and examples of the
functions, or other articles. More
articles will be added in the future for other scenarios.
Monte Carlo Confidence Intervals
Monte Carlo confidence intervals can also be formed using the
functions illustrated above. First use do_mc() instead of
do_boot() to generate simulated sample estimates. When
calling other main functions, use mc_ci = TRUE and set
mc_out to the output of do_mc(). Please refer
to vignette("do_mc") for an illustration, and
vignette("do_mc_lavaan_mi") on how to form Monte Carlo
confidence intervals for models fitted to multiple imputation
datasets.
References
Cheung, M. W.-L. (2009). Comparison of methods for constructing confidence intervals of standardized indirect effects. Behavior Research Methods, 41(2), 425-438. https://doi.org/10.3758/BRM.41.2.425
Cheung, S. F., & Cheung, S.-H. (2024). manymome: An R package for computing the indirect effects, conditional effects, and conditional indirect effects, standardized or unstandardized, and their bootstrap confidence intervals, in many (though not all) models. Behavior Research Methods, 56(5), 4862–4882. https://doi.org/10.3758/s13428-023-02224-z
Falk, C. F., & Biesanz, J. C. (2015). Inference and interval estimation methods for indirect effects with latent variable models. Structural Equation Modeling: A Multidisciplinary Journal, 22(1), 24–38. https://doi.org/10.1080/10705511.2014.935266
Friedrich, R. J. (1982). In defense of multiplicative terms in multiple regression equations. American Journal of Political Science, 26(4), 797-833. https://doi.org/10.2307/2110973
Hayes, A. F. (2015). An index and test of linear moderated mediation. Multivariate Behavioral Research, 50(1), 1-22. https://doi.org/10.1080/00273171.2014.962683
Hayes, A. F. (2018). Partial, conditional, and moderated moderated mediation: Quantification, inference, and interpretation. Communication Monographs, 85(1), 4-40. https://doi.org/10.1080/03637751.2017.1352100
Hayes, A. F. (2022). Introduction to mediation, moderation, and conditional process analysis: A regression-based approach (Third Edition). The Guilford Press.
Kwan, J. L. Y., & Chan, W. (2018). Variable system: An alternative approach for the analysis of mediated moderation. Psychological Methods, 23(2), 262-277. https://doi.org/10.1037/met0000160
Miles, J. N. V., Kulesza, M., Ewing, B., Shih, R. A., Tucker, J. S., & D’Amico, E. J. (2015). Moderated mediation analysis: An illustration using the association of gender with delinquency and mental health. Journal of Criminal Psychology, 5(2), 99-123. https://doi.org/10.1108/JCP-02-2015-0010
Tofighi, D., & Kelley, K. (2020). Indirect effects in sequential mediation models: Evaluating methods for hypothesis testing and confidence interval formation. Multivariate Behavioral Research, 55(2), 188–210. https://doi.org/10.1080/00273171.2019.1618545