Generate Monte Carlo Estimates
Shu Fai Cheung & Sing-Hang Cheung
2026-03-26
Source:vignettes/do_mc.Rmd
do_mc.RmdIntroduction
This article is a brief illustration of how to use
do_mc() from the package manymome (Cheung & Cheung,
2024) to generate Monte Carlo estimates for
indirect_effect() and cond_indirect_effects()
to form Monte Carlo confidence intervals.
Although indirect_effect() and
cond_indirect_effects() can also be used to generate Monte
Carlo estimates when they are called (see
vignette("manymome")), there may be situations in which
users prefer generating the Monte Carlo estimates first before calling
indirect_effect() and cond_indirect_effects().
do_mc() is for this purpose.
Monte Carlo confidence intervals only support models fitted by
lavaan::sem() and (since version 0.2.7.3)
lavaan.mi::sem.mi() or lavaan.mi::lavaan.mi().
(Support for semTools::runMI() and its wrappers was removed
in version 0.2.7.3 because this and related functions were deprecated in
semTools.)
How It Works
The function do_mc() retrieves the variance-covariance
matrix of the parameter estimates and then generates a number of sets of
simulated sample estimates using a multivariate normal distribution.
Other parameters and implied variances, covariances, and means of
variables are then generated from these simulated estimates.
When a % Monte Carlo confidence interval is requested, the th percentile and the th percentile are used to form the confidence interval. For a 95% Monte Carlo confidence interval, the 2.5th percentile and 97.5th percentile will be used.
The Workflow
The following workflow will be demonstrated;
Fit the model as usual.
Use
do_mc()to generate the Monte Carlo estimates.Call other functions (e.g,
indirect_effect()andcond_indirect_effects()) to compute the desired effects and form Monte Carlo confidence intervals.
Demonstration
Fit a Model by lavaan::sem()
The data set for illustration:
library(manymome)
dat <- data_med
head(dat)
#> x m y c1 c2
#> 1 9.931992 17.89644 20.73893 1.426513 6.103290
#> 2 8.331493 17.92150 22.91594 2.940388 3.832698
#> 3 10.327471 17.83178 22.14201 3.012678 5.770532
#> 4 11.196969 20.01750 25.05038 3.120056 4.654931
#> 5 11.887811 22.08645 28.47312 4.440018 3.959033
#> 6 8.198297 16.95198 20.73549 2.495083 3.763712It has one predictor (x), one mediator (m),
one outcome variable (y), and two control variables
(c1 and c2).
The following simple mediation model with two control variables
(c1 and c2) will be fitted:
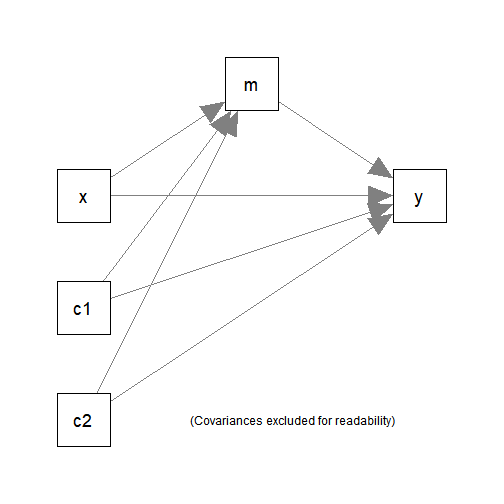
Fit the model by lavaan::sem():
mod <-
"
m ~ x + c1 + c2
y ~ m + x + c1 + c2
"
fit_lavaan <- sem(model = mod, data = dat,
fixed.x = FALSE,
estimator = "MLR")
summary(fit_lavaan)
#> lavaan 0.6-21 ended normally after 1 iteration
#>
#> Estimator ML
#> Optimization method NLMINB
#> Number of model parameters 15
#>
#> Number of observations 100
#>
#> Model Test User Model:
#> Standard Scaled
#> Test Statistic 0.000 0.000
#> Degrees of freedom 0 0
#>
#> Parameter Estimates:
#>
#> Standard errors Sandwich
#> Information bread Observed
#> Observed information based on Hessian
#>
#> Regressions:
#> Estimate Std.Err z-value P(>|z|)
#> m ~
#> x 0.935 0.075 12.437 0.000
#> c1 0.198 0.079 2.507 0.012
#> c2 -0.168 0.099 -1.703 0.089
#> y ~
#> m 0.785 0.233 3.363 0.001
#> x 0.508 0.323 1.573 0.116
#> c1 0.140 0.188 0.747 0.455
#> c2 -0.154 0.214 -0.720 0.471
#>
#> Covariances:
#> Estimate Std.Err z-value P(>|z|)
#> x ~~
#> c1 0.026 0.121 0.211 0.833
#> c2 0.100 0.084 1.186 0.235
#> c1 ~~
#> c2 -0.092 0.109 -0.841 0.400
#>
#> Variances:
#> Estimate Std.Err z-value P(>|z|)
#> .m 0.681 0.085 7.976 0.000
#> .y 4.030 0.580 6.944 0.000
#> x 1.102 0.150 7.338 0.000
#> c1 1.218 0.161 7.540 0.000
#> c2 0.685 0.073 9.340 0.000Suppose we would like to use robust “sandwich” standard errors and
confidence intervals provided by MLR for the free parameters, but want
to use Monte Carlo confidence interval for the indirect effect. In the
call above, we used estimator = "MLR" and did not set
se = "boot".
Generate Monte Carlo Estimates
We can then call do_mc() on the output of
lavaan::sem() to generate the Monte Carlo estimates of all
free parameters and the implied statistics, such as the
variances of m and y, which are not free
parameters but are needed to form the confidence interval of the
standardized indirect effect.
mc_out_lavaan <- do_mc(fit = fit_lavaan,
R = 10000,
seed = 4234)
#> Stage 1: Simulate estimates
#> Stage 2: Compute implied statisticsUsually, just three arguments are needed:
fit: The output oflavaan::sem().R: The number of Monte Carlo replications. Should be at least 10000 in real research.seed: The seed for the random number generator. To be used byset.seed(). It is recommended to set this argument such that the results are reproducible.
Parallel processing is not used. However, the time taken is rarely long because there is no need to refit the model many times.
Using the Output of do_mc() in Other Functions of
manymome
When calling indirect_effect() or
cond_indirect_effects(), the argument mc_out
can be assigned the output of do_mc(). They will then
retrieve the stored simulated estimates to form the Monte Carlo
confidence intervals, if requested.
out_lavaan <- indirect_effect(x = "x",
y = "y",
m = "m",
fit = fit_lavaan,
mc_ci = TRUE,
mc_out = mc_out_lavaan)
out_lavaan
#>
#> == Indirect Effect ==
#>
#> Path: x -> m -> y
#> Indirect Effect: 0.733
#> 95.0% Monte Carlo CI: [0.306 to 1.174]
#>
#> Computation Formula:
#> (b.m~x)*(b.y~m)
#>
#> Computation:
#> (0.93469)*(0.78469)
#>
#>
#> Monte Carlo confidence interval with 10000 replications.
#>
#> Coefficients of Component Paths:
#> Path Coefficient
#> m~x 0.935
#> y~m 0.785Reusing the simulated estimates can ensure that all analyses with Monte Carlo confidence intervals are based on the same set of simulated estimates, without the need to generate these estimates again.
Missing Data
Monte Carlo confidence intervals can be formed when the
variance-covariance matrix of the parameter estimates can be retrieved.
Therefore, do_mc() can be used when missing data is handled
by full information maximum likelihood in lavaan using
missing = "fiml". It also supports multiple imputation if
lavaan.mi::sem.mi() or lavaan.mi::lavaan.mi()
(since version 0.2.7.3). See vignette("do_mc_lavaan_mi")
for an illustration.
The Structure of the Output
The output of do_mc() in this case is an object of the
class mc_out, which is a list of R lists, each
with two elements: est and implied_stats.
This is the content of est of the first list:
mc_out_lavaan[[1]]$est
#> lhs op rhs est
#> 1 m ~ x 0.860
#> 2 m ~ c1 0.160
#> 3 m ~ c2 -0.086
#> 4 y ~ m 0.567
#> 5 y ~ x 0.881
#> 6 y ~ c1 0.008
#> 7 y ~ c2 0.117
#> 8 m ~~ m 0.619
#> 9 y ~~ y 3.431
#> 10 x ~~ x 1.061
#> 11 x ~~ c1 -0.113
#> 12 x ~~ c2 0.099
#> 13 c1 ~~ c1 1.190
#> 14 c1 ~~ c2 -0.243
#> 15 c2 ~~ c2 0.724The content is just the first four columns of the output of
lavaan::parameterEstimates(). Note that only fixed and free
parameters are used so other rows, if any, are not used even if
present.
This is the content of implied_stats of the first
list:
mc_out_lavaan[[1]]$implied_stats
#> $cov
#> m y x c1 c2
#> m 1.401
#> y 1.575 5.619
#> x 0.886 1.448 1.061
#> c1 0.114 -0.053 -0.113 1.190
#> c2 -0.016 0.161 0.099 -0.243 0.724
#>
#> $mean
#> m y x c1 c2
#> NA NA NA NA NAIt has three elements. cov is the implied variances and
covariances of all variables. If a model has latent variables, they will
be included too. The other elements, mean and
mean_lv, are the implied means of the observed variables
and the latent variables (if any), respectively. The elements are
NA if mean structure is not in the fitted model.
Limitations
Monte Carlo confidence intervals require the variance-covariance
matrix of all free parameters. Therefore, only models fitted by
lavaan::sem() and (since 0.2.7.3)
lavaan.mi::sem.mi() or lavaan.mi::lavaan.mi()
are supported. Models fitted by stats::lm() do not have a
variance-covariance matrix for the regression coefficients from two or
more regression models and so are not supported by
do_mc().
Further Information
For further information on do_mc(), please refer to its
help page.
Reference
Cheung, S. F., & Cheung, S.-H. (2024). manymome: An R package for computing the indirect effects, conditional effects, and conditional indirect effects, standardized or unstandardized, and their bootstrap confidence intervals, in many (though not all) models. Behavior Research Methods, 56(5), 4862–4882. https://doi.org/10.3758/s13428-023-02224-z